Per gene statistical comparison of CVs: permutation-based p-values
Joyce Hsiao
2015-11-17
Last updated: 2015-12-11
Code version: 7a25b2aef5c8329d4b5394cce595bbfc2b5b0c3e
Objective
Quantify statistical significance of individual differences between adjusted CVs.
We permuted the wells labels across samples for 10,000 times. Then, for each gene, we computed the number of permuted samples produced larger distance than the observed distance between standardized coefficients of variance between individuals.
Set up
library("data.table")
library("dplyr")
library("limma")
library("edgeR")
library("ggplot2")
library("grid")
library("Humanzee")
theme_set(theme_bw(base_size = 12))
source("functions.R")Prepare data
Input annotation of only QC-filtered single cells. Remove NA19098.r2
anno_filter <- read.table("../data/annotation-filter.txt",
header = TRUE,
stringsAsFactors = FALSE)
dim(anno_filter)[1] 568 5Import endogeneous gene molecule counts that are QC-filtered, CPM-normalized, ERCC-normalized, and also processed to remove unwanted variation from batch effet. ERCC genes are removed from this file.
molecules_ENSG <- read.table("../data/molecules-final.txt",
header = TRUE, stringsAsFactors = FALSE)
stopifnot(NCOL(molecules_ENSG) == NROW(anno_filter))Compute normalized CV
We compute squared CV across cells for each individual and then for each individual CV profile, account for mean dependency by computing distance with respect to the data-wide coefficient variation on the log10 scale.
ENSG_cv <- Humanzee::compute_cv(log2counts = molecules_ENSG,
grouping_vector = anno_filter$individual)
ENSG_cv_adj <- Humanzee::normalize_cv(group_cv = ENSG_cv,
log2counts = molecules_ENSG,
anno = anno_filter)Compute summary measure of deviation
- Standardize the each CV vectors
Individual CV vectors are standarized for individual CV mean and coefficients of variation across genes.
df_cv <- data.frame(NA19098 = ENSG_cv_adj[[1]]$log10cv2_adj,
NA19101 = ENSG_cv_adj[[2]]$log10cv2_adj,
NA19239 = ENSG_cv_adj[[3]]$log10cv2_adj)
library(matrixStats)
df_norm <- sweep(df_cv, MARGIN = 2, STATS = colMeans(as.matrix(df_cv)), FUN = "-")
df_norm <- sweep(df_norm, MARGIN = 2, STATS = sqrt(colVars(as.matrix(df_cv))), FUN = "/")
colnames(df_norm) <- names(ENSG_cv_adj)- SSM and SAM
Compute metrics for quantifying similarity between the three individual coefficients of variation.
library(matrixStats)
df_norm <- as.data.frame(df_norm)
df_norm$squared_dev <- rowSums( ( df_norm - rowMedians(as.matrix(df_norm)) )^2 )
df_norm$abs_dev <- rowSums(abs( df_norm - rowMedians(as.matrix(df_norm)) ))Compute permuted p-vals
Export some rdas.
if (!file.exists("rda/cv-adjusted-statistical-test-permute/adj-cv.rda")) {
save(df_norm, file = "rda/cv-adjusted-statistical-test-permute/adj-cv.rda")
}
if (!file.exists("rda/cv-adjusted-statistical-test-permute/permute-cv-test.rda")) {
save(molecules_ENSG, anno_filter,
file = "rda/cv-adjusted-statistical-test-permute/permute-cv-test.rda")
}Compute permuted distances for each gene.
sbatch ../code/permuted-cv-test.sbatchCompute permutation-based p-values for each gene.
sbatch ../code/permuted-cv-compute-pval.sbatchSanity checks on the permuted p-values
Confirm that the gene with the larger SSM and SAM are significant with a p-value of 0.
load("rda/cv-adjusted-statistical-test-permute/permuted-pval.rda")
permuted_pval$squared_dev[which.max(df_norm$squared_dev)][1] 0permuted_pval$squared_dev[order(df_norm$squared_dev, decreasing = T)[3]][1] 0library(ggplot2)
ggplot(data.frame(pvals = permuted_pval$squared_dev,
dist = df_norm$squared_dev),
aes(x = dist, y = pvals)) +
geom_point()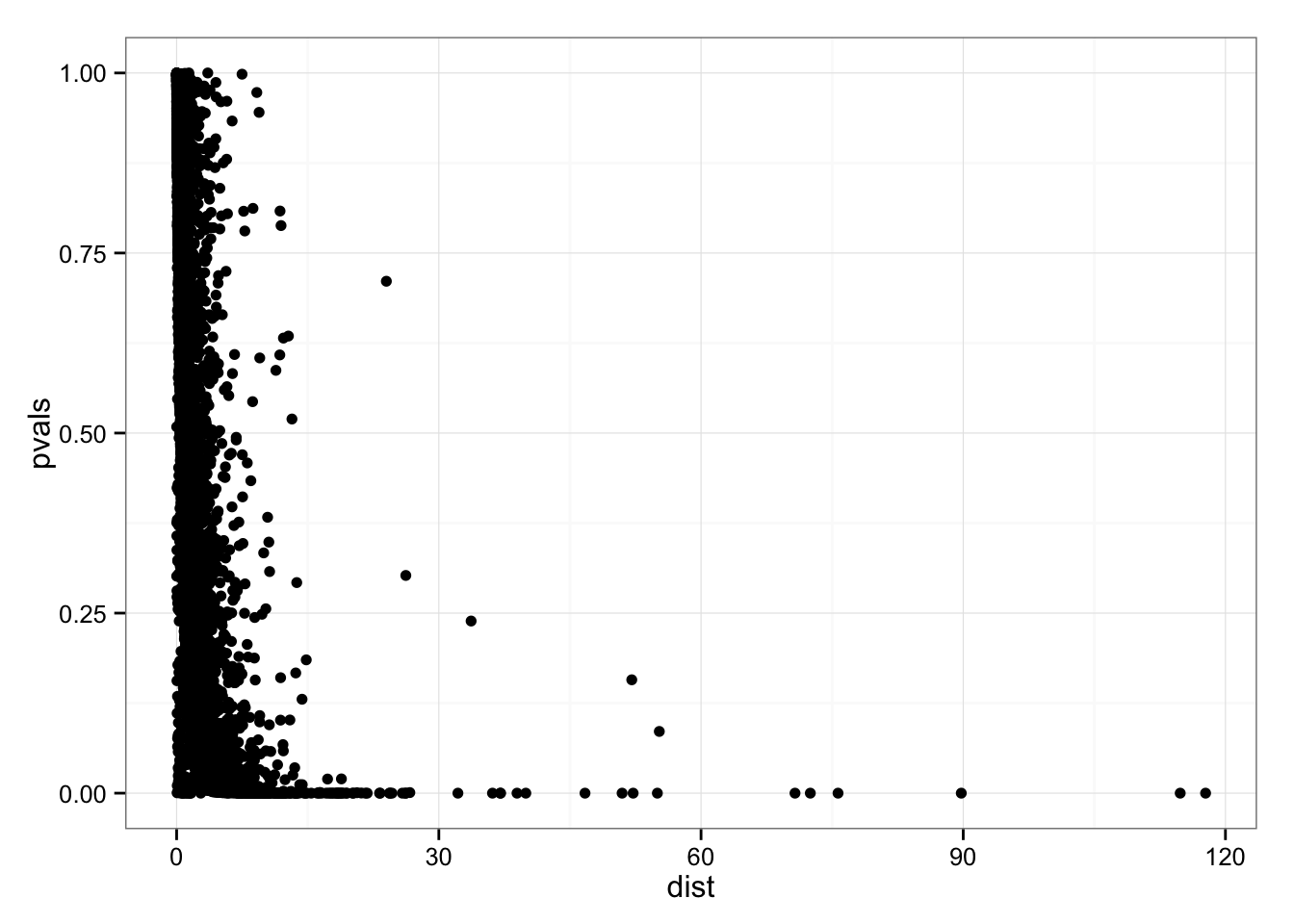
Gene with the largest Sum-of-Squared-Deviations.
ggplot(do.call(rbind, ENSG_cv_adj),
aes(x = log10(mean), y = log10cv2_adj)) +
geom_point(aes(col = group), cex = .8) + facet_wrap( ~ group) +
ggtitle("Gene with the largest SSM") +
geom_point(data = subset(do.call(rbind, ENSG_cv_adj),
c(1:NROW(ENSG_cv_adj[[1]])) %in%
order(df_norm$squared_dev, decreasing = TRUE )[1]),
colour = "grey20")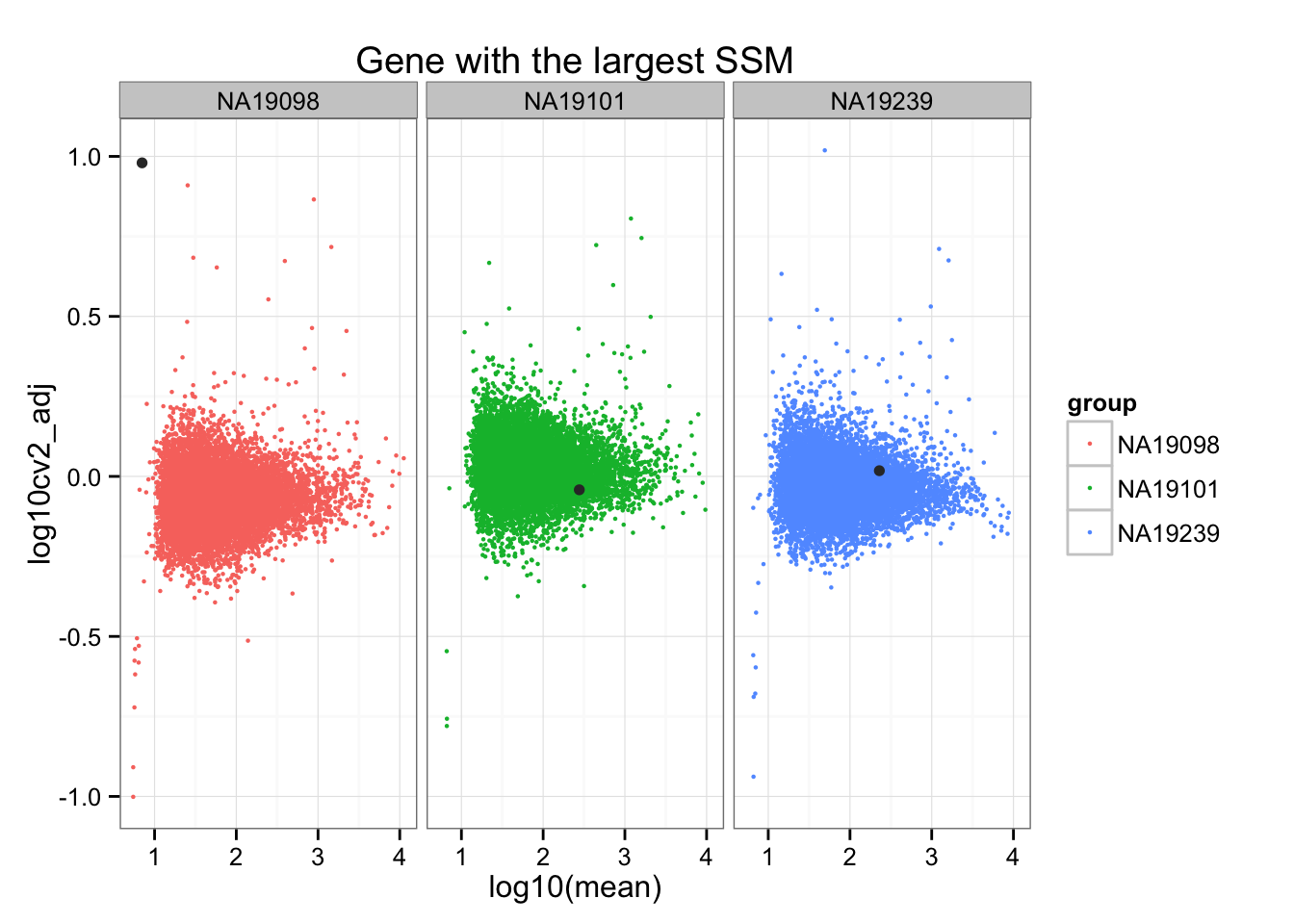
Session information
sessionInfo()R version 3.2.1 (2015-06-18)
Platform: x86_64-apple-darwin13.4.0 (64-bit)
Running under: OS X 10.10.5 (Yosemite)
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] grid stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] matrixStats_0.15.0 zoo_1.7-12 Humanzee_0.1.0
[4] ggplot2_1.0.1 edgeR_3.10.5 limma_3.24.15
[7] dplyr_0.4.3 data.table_1.9.6 knitr_1.11
loaded via a namespace (and not attached):
[1] Rcpp_0.12.2 magrittr_1.5 MASS_7.3-45 munsell_0.4.2
[5] lattice_0.20-33 colorspace_1.2-6 R6_2.1.1 stringr_1.0.0
[9] plyr_1.8.3 tools_3.2.1 parallel_3.2.1 gtable_0.1.2
[13] DBI_0.3.1 htmltools_0.2.6 yaml_2.1.13 assertthat_0.1
[17] digest_0.6.8 reshape2_1.4.1 formatR_1.2.1 evaluate_0.8
[21] rmarkdown_0.8.1 labeling_0.3 stringi_1.0-1 scales_0.3.0
[25] chron_2.3-47 proto_0.3-10